Edaphobase, a non-commercial data infrastructure developed by the Senckenberg Museum of Natural History Görlitz in Germany, combines data from heterogeneous sources on soil animals, their distribution and habitat parameters of their sites of occurrence and makes these data available to the public (open access).
The data in Edaphobase originate from the scientific literature, unpublished results of field studies (theses, reports), collections of museums and research institutions as well as raw data from research studies and well-founded observations. Data types comprise modern taxonomic nomenclatures and synonyms, geographical references, quantities of collected organisms, soil parameters, vegetation, meteorological data, sampling and extraction methods, identification methods, preparation techniques and behavioural data. Altogether, data can be entered into (currently) ca. 620 available data fields in the above-mentioned categories.
Edaphobase currently includes data on Nematoda, Collembola, Oribatida, Gamasina, Chilopoda, Diplopoda, Isopoda, Enchytraeidae, and Lumbricidae. The data model allows adding additional taxonomic groups easily, as long as the current (nomenclaturally complete) taxonomy and systematics of the group are provided. Edaphobase combines data on the taxonomy, zoogeography and ecology of these organisms in a comprehensive manner.
Edaphobase is strongly geared towards common data re-use and provision of data and online analysis tools. It follows the FAIR principles and can offer DOIs for submitted data sets. Metadata are defined (including mandatory and recommended fields; in line with i.e. DataCite and INSPIRE standards), as are all data fields in the data base (including formats, units, etc.).
The database is publically available (open access) via a web-based data-query browser portal. All data (provided it is not anonymized or in an embargo period, see below) can be queried and collated via multiple filters and be downloaded by registered users. Simple queries are possible as well as more sophisticated analyses of different data groups. Specific applications include, e.g.,the elucidation of species-specific habitat preferences (niche space), distribution patterns or environmental correlations with population densities.
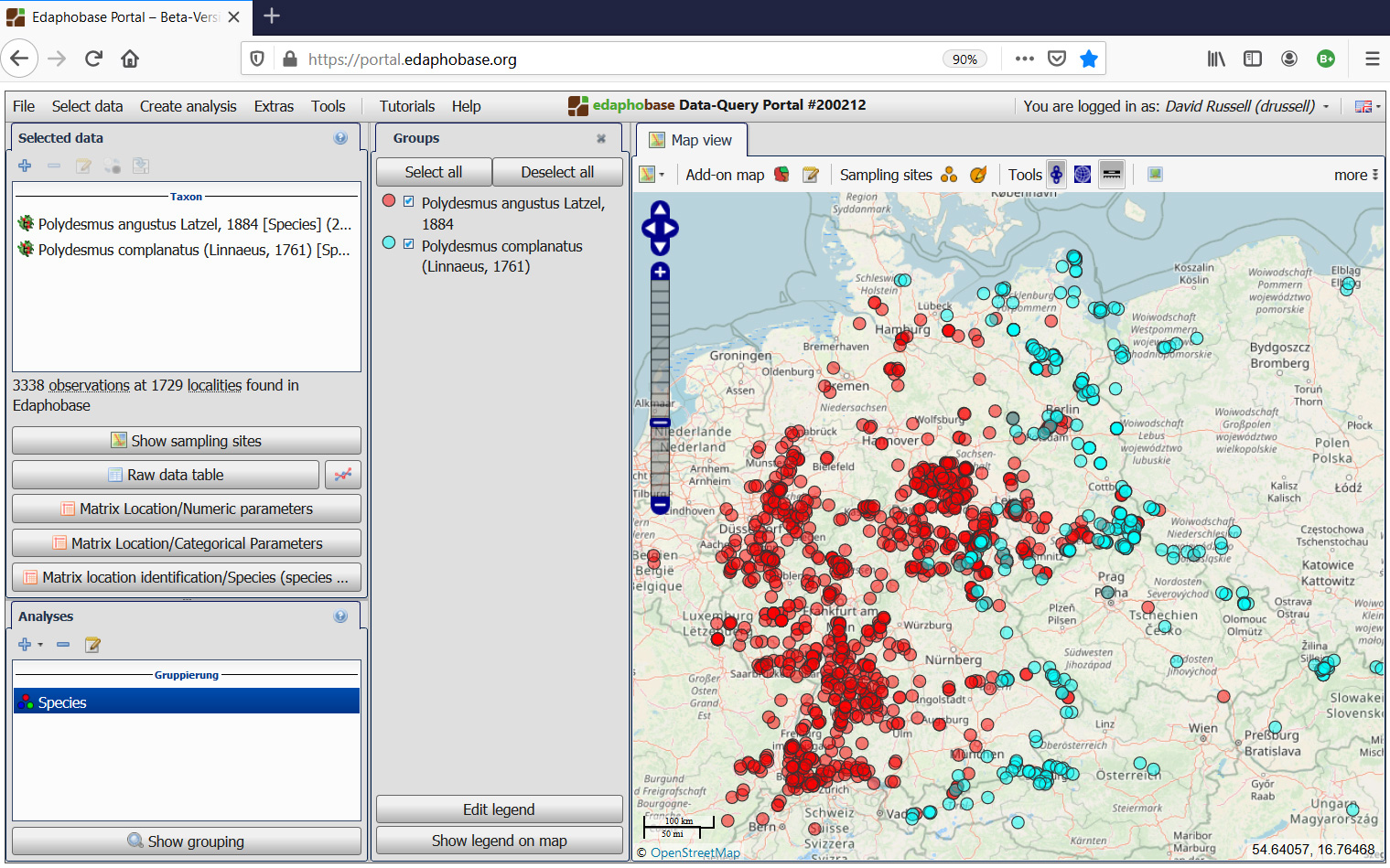
The Query Portal offers detailed query filters and allows collation of database data into one file for analysis or data download, whereby the specific data fields in the table can be specified by the user. Almost all data fields are available for download. Sites of occurrence for all taxa can be mapped in the browser application, differentiated according to species, habitat types, soil parameters etc. Basic descriptive analysis tools regarding niche space of species as well as expected species composition for site-parameter combinations are available. More detailed prognoses of soil biodiversity (maps as well as point scale for specific sites) are currently being developed within the EUdaphobase COST Action as well as other cooperative projects.
Data import software allows available data to be imported “as is” (from text, EXCEL or ACCESS files) and mapped to Edaphobase (including semantic harmonization). Thesauri for various variable/parameter labels are being compiled via the software.
Data-quality is ensured primarily during data-import procedures via automatic controls in the import software (pre-import quality control), a standardized quality-control checklist between data upload and final import in the database (peri-import quality control) and well as control procedures for the data provider in the web-browser application after data is imported (post-import control). This includes qualitative control procedures for taxonomic data, whereby an international review board of recognized taxonomic experts is being developed by the EUdaphobase COST Action.
Edaphobase has a legally reviewed data policy that is in line with the recent EU and German data-protection legislation, outlines how uploaded data is publically provided in a browser-based web application while protecting IPRs, offers possibilities for public data anonymization and/or an embargo period, as well a data-sharing agreement document.
Data from Edaphobase is regularly uploaded to the Global Biodiversity Information Facility (GBIF) network.
- Further information can be found here
- Explore the database here
- Submit data